Ambitious Google drive to put human genome online gathers steam
Published 8 Nov, 2014 15:44 | Updated 8 Nov, 2014 16:12
Google’s plan to store entire copies of the human genome online is edging closer to reality. With 3,500 genomes already stored on its servers and more medical institutes jumping onboard, the blueprint of every person on Earth could soon be in the cloud.
The potentially game-changing project, called Google Genomics, has now been quietly moving forward for a year and a half.
Perhaps with so many ambitious plans coming out of the company’s secretive Google X research and development division, from nanobots to sniff out cancer to tremor-canceling spoons for Parkinson's patients, it’s easy for even deeply ambitious projects to get overlooked.
READ MORE: Google’s next data collection project: Human body
RT first wrote about the search giant’s plan to create individual genome databases in July, but even promotional videos hashing out the details of the project attracted just 5,000 views over the past four months.
Maybe the fact that Google Drive currently cannot cope with entire copies of the genome has left many thinking the project is mere speculative pipe dream at this stage of the game.
READ MORE: Google nanobots: Early warning system for cancer, heart disease inside the body
But for futurists, whose first commandment is Moore's Law, that which is impossible today will likely be outdated tomorrow.
Google itself was quick to point out that at the inception of the human Genome project, it took 15 years and $3 billion just to do the first human genome sequence. Today, it can all be done in a day, and for about $1,000.
Just how many gigs am I?
But just how much memory is needed to save all 6 billion of the nucleotide letters that comprise a single genome sequence? Google estimates it’s around 100 gigabits, which might not seem like a lot, until you consider just how many of us there are.
For example, if you wanted to read the DNA of everyone (officially!) living in Moscow, it would take more than 1.2 million terabit hardrives. While that is obviously an enormous amount of information to process, Google’s current search index stands at 100 petabyes – 100,000 terabytes. The average search query, however, takes 0.25 seconds.
And it is applying this self-same search technology to the Google Genomics which is viewed as the key.
At the inception of the project, scientists began hammering out an application programming interface (API) which would allow them to move DNA data into Google server clusters and conduct experiments using the companies renowned web-indexing technology.
And as scientists have expanded their studies beyond individual genomes, hammering out a synthesis between data science and life science could propel the pace of medical advancement over the coming years.
“We saw biologists moving from studying one genome at a time to studying millions,” David Glazer, the software engineer who led the effort and was previously head of platform engineering for Google+, the social network, told the MIT Technology Review. “The opportunity is how to apply breakthroughs in data technology to help with this transition.”
Currently, different genome data sets are exclusively available to specific research labs. The goal then, is to create one centralized database where researchers can compare millions of genome sequences at one time.
Speaking to Technology Review, Sheila Reynolds, a research scientist at the Institute for Systems Biology in Seattle, one idea is to create “cancer genome clouds” where scientists can share information and quickly run virtual experiments as easily as a web search.
“Our bird’s eye view is that if I were to get lung cancer in the future, doctors are going to sequence my genome and my tumor’s genome, and then query them against a database of 50 million other genomes,” Deniz Kural, CEO of Seven Bridges, which stores genome data on behalf of 1,600 researchers in Amazon’s cloud, told the magazine. “The result will be ‘Hey, here’s the drug that will work best for you.’”
But as Reynolds noted, not every research institute has the ability to download a petabyte of data, or the computing power to analyze it.
With a centralized database, however, those technological trammels would be put out to pasture.
The treatment potential of being able to compare the genomes of multiple individuals suffering from the same ailments is astronomical, as is the profit motive for whoever holds the keys to the data locker.
This reality has already put Google, Amazon and Microsoft and IBM in a race to see who will store the data. And on a fair playing field, the competition has driven prices down.

Saving you for a quarter a year
Currently, storing a single human genome with Google is going to cost you $25 a year, in the same ballpark as Amazon. Running analysis of the data, of course, is gonna cost you. The catch, of course, is that people’s DNA is 99.1 percent identical. Once you can whittle it down to the 0.1 percent that makes us who we are, less than a gig will be needed to store the essence of you in the cloud. So in the long term, a bit of analysis and a quarter will get your unique genomic sequence put up in the cloud for a year.
Glazer did tell the magazine just how many customers Google Genomics has now, though at least 3,500 genomes from public projects are already stored on Google’s server farm.
According to The Verge, the National Cancer Institute has already signed on to the project, and has expressed its willingness to pay $19 million to upload copies of its 2,600 terabyte Cancer Genome Atlas to Google Genomics and Amazon’s data center.
The project, however, definitely comes with its privacy pitfalls.
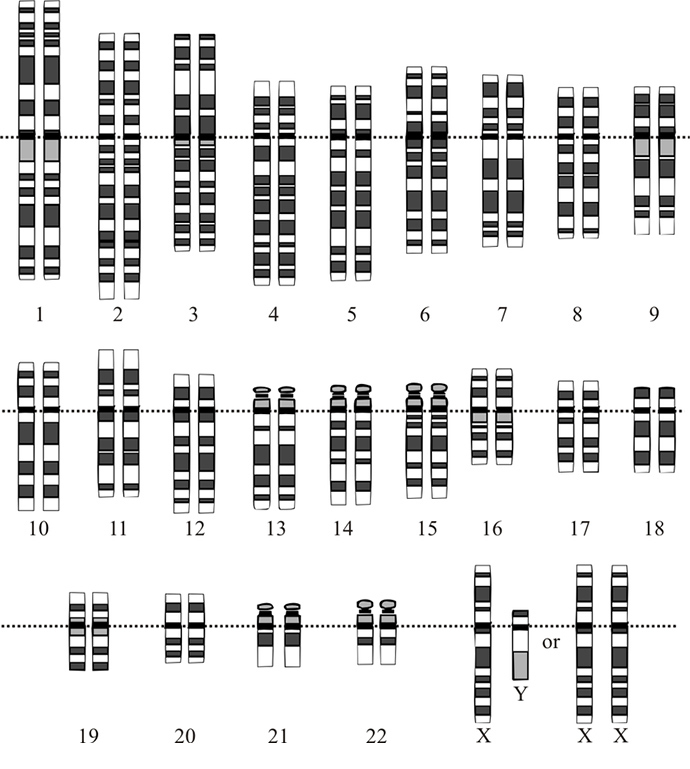
As Gizmodo recently noted, a study in the Journal Science last year showed it was possible to identify several men from the publicly available 1000 Genomes Project based on their Y chromosomes and age, location, and family tree data.
Insurance companies would also likely be thrilled to get their hands on that data.
There is also the issue of whether scientists should tell people if they unknowingly have a rare disease, or have unknown siblings out there in the world.
But while both concerns of privacy and practicality are inevitable in any venture of this scope, the likelihood that the seemingly infinite permutations of AGCT which tell the story of every person on earth seems all but inevitable.















