New types of human brain cells found in quest for understanding its development & dysfunctions
Published 19 Aug, 2017 23:36 | Updated 20 Aug, 2017 09:26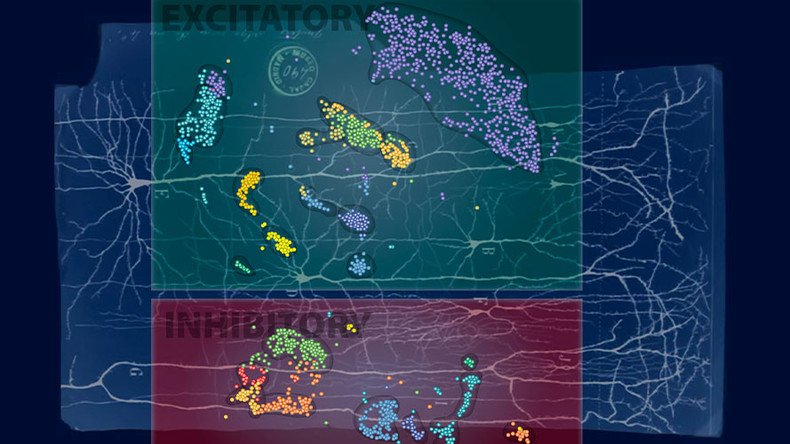
Seeking to find out how the human brain works on a molecular level, scientists used a new method of categorizing brain cells and found new subtypes, which promises to significantly increase understanding of how a brain develops and why things go wrong.
A team of researchers from the Salk Institute in California began their work by examining cells from the frontal cortices of mice and humans.
The frontal cortex is the area of the brain responsible for some of an individual’s most important characteristics including personality, decision making and complex thinking.
The team then compiled incredibly detailed information on what makes one individual brain cell different from another. The cells can appear structurally identical to each other yet still behave differently.
READ MORE: DNA privacy protection tackled by new encryption technology – study
“We think it’s pretty striking that we can tease apart a brain into individual cells, sequence their methylomes, and identify many new cell types along with their gene regulatory elements, the genetic switches that make these neurons distinct from each other, ” one of the authors of the study, professor Joseph Ecker, explained.
The team was trying to identify how many types of neurons exist. This is one of the great unanswered questions in neuroscience and experts believe it could lead to a dramatically better understanding of brain development and dysfunction.
“This opens up the possibility of understanding what makes two neurons – that sit in the same brain region and otherwise look similar – behave differently,” said Margarita Behrens, co-senior author on the paper.
“This is a very novel study both technically and conceptually,” explained Hongkui Zeng of the University of Seattle to The San Diego Tribune.
“Just like each person is unique, each cell or neuron is also different from all other cells in many ways. Thus cell type classification is a very challenging task, one needs to profile a cell from multiple aspects and combine all the information together in order to assign cell type identity correctly.”
Researchers found that the neurons from the human brain were more diverse, forming 21 subtypes to the mice’s 16. They also identified unique human neuron subtypes that had never been defined before.
“These results open the door to a deeper understanding of what sets human brains apart from those of other animals,” the researchers said in a statement.
The study was published in the journal Science. The researchers plan to expand the study to look at more parts of the brain.
The team say their next goal is to create a “parts list” of both mouse and human brains. After that is complete, they plan to compare the patterns of gene activation in healthy people with those who suffer from brain diseases.










